The William Tong Prize 2024 – Unveiling the unknown: metagenomics to diagnose unexpected causes of encephalitis
The William Tong prize was established in 2018 in memory of the late William Tong, who had just stepped down as Chair of the College Panel of Examiners in Virology. William Tong’s family and friends raised funds to establish the prize in his name, to be awarded to a trainee or student for outstanding original work in clinical virology.
At this year’s UK Clinical Virology Network annual conference, the William Tong Prize was awarded to Dr Julianne Brown for her work on metagenomics in diagnosing unexpected causes of encephalitis.
Dr Julianne Brown is a Higher Specialist Scientist Training (HSST) trainee and Principal Clinical Scientist in Molecular Microbiology and Virology at Great Ormond Street Hospital for Children NHS Foundation Trust, where she is responsible for delivering the clinical molecular microbiology service for diagnosis of infection, including bacterial and fungal polymerase chain reaction (PCR) and untargeted metagenomics for pathogen detection in sterile site and respiratory specimens.
Case outline
A 16-year-old boy presented to hospital with progressive headache, ascending muscle weakness, generalised tonic-clonic seizures and myoclonic jerks. He was disorientated with photophobia, phonophobia and difficulty swallowing and chewing. Progressive breathing difficulties due to weakness led to ventilation and admission to an intensive care unit.15 years earlier he had undergone a haematopoetic stem cell transplant (HSCT) following a diagnosis of primary hemophagocytic lymphohistiocystosis (HLH). At 10 years old, he was started on immunoglobulin replacement therapy for antibody deficiency but was otherwise previously well, cognitively normal and attending school with no neurological concerns.
On admission he was found to be severely lymphopenic and brain magnetic resonance imaging (MRI) showed bilateral widespread cortical changes. A brain biopsy and 3 cerebrospinal fluids (CSFs) were all negative by culture and by PCR targeting several viruses and bacteria. Clinical and radiological findings were suggestive of central nervous system HLH; therefore he was treated with intensive immunosuppression, but with no response.
Metagenomics was performed on the brain biopsy. Unlike PCR, which targets and detects specific genome sequences, metagenomics sequences all the RNA and DNA in a sample which includes not only human sequences but also the genomes of any organisms present. The human sequences are removed, and the remaining sequences are analysed to identify any organism present. Since the method is untargeted it allows detection of rare, unexpected or novel organisms (Figure 1).
Metagenomics uncovered the presence of avian paramyxovirus 1 (APMV-1), later confirmed in brain tissue by PCR and immunohistochemistry. APMV-1 is shed in pigeon faeces and is endemic worldwide. In birds it can be highly pathogenic, causing pneumoencephalitis (also known as Newcastle Disease) with up to 100% mortality. Notably the virus was not detectable in CSF by any method, therefore the diagnosis was only made possible with brain biopsy. Following the APMV-1 diagnosis, the patient’s immunosuppressive medication was reduced and he was treated with antivirals. However, he continued to deteriorate and unfortunately died 8 months after symptom onset.
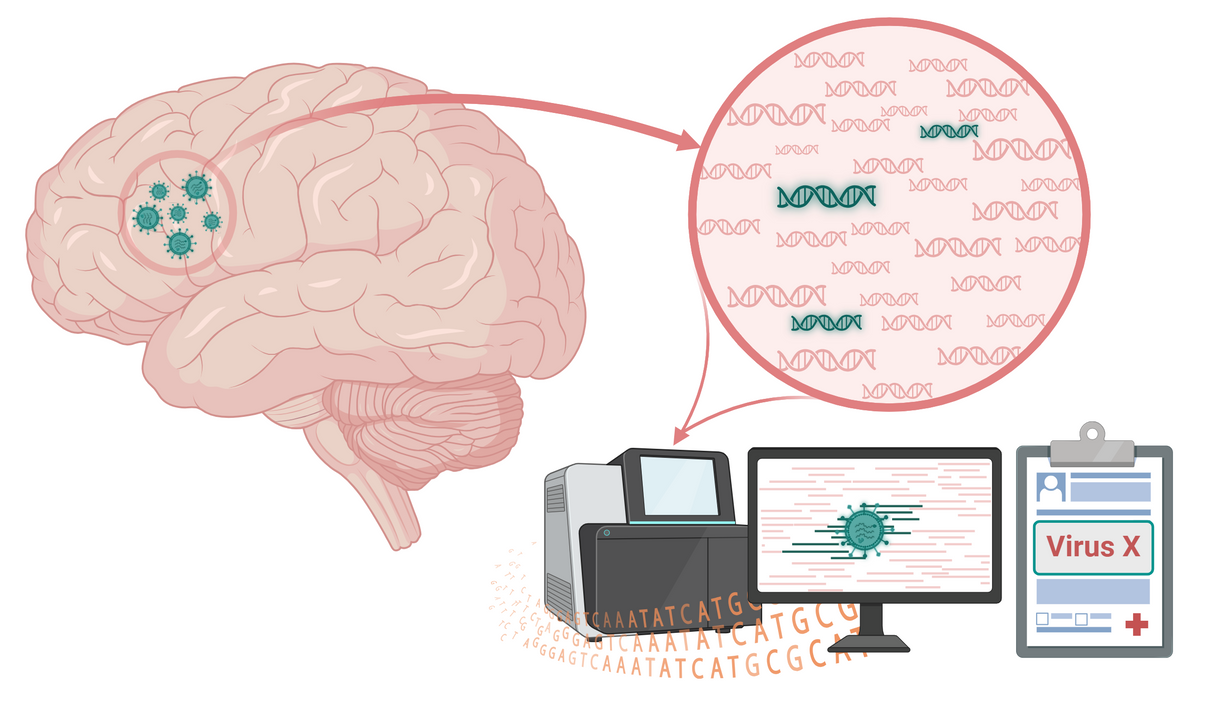
Figure 1: Schematic of clinical metagenomics to diagnose infectious causes of encephalitis. Total DNA and RNA are sequenced from a brain biopsy (or CSF), including human and non-human genomes. Non-human sequences are analysed to identify any organisms present in the specimen. Metagenomics is untargeted and therefore requires no a priori assumptions or knowledge of the causative organism.
Metagenomics in encephalitis diagnosis
Despite being endemic in birds, APMV-1 is an extremely rare cause of severe disease in humans. Only 2 other cases of fatal encephalitis caused by APMV-1 have been described worldwide, both in immunosuppressed children; 1 in France and the other in Australia.1,2 For this reason, APMV-1 is not part of targeted testing in encephalitis. Diagnosis of infectious encephalitis largely relies on culture and/or targeted PCR in CSFs and brain biopsies. This works well for infections with well-characterised aetiologies, in which all the causative organisms are known and can be tested for.
However infectious encephalitis has an extremely broad aetiology. The list of causative organisms becomes even longer if a patient is immunosuppressed or has a relevant travel history. CSFs and brain biopsies are small-volume, precious samples. Therefore, testing is usually limited to the most common organisms associated with disease; if they are negative, the cause remains undiagnosed. Consequently, for a third of cases of encephalitis there is no diagnosis.3
Metagenomics is proving to be instrumental in the diagnosis of infectious encephalitis caused by rare, unusual or unexpected organisms that are unlikely to be or cannot be detected by other methods. At the Great Ormond Street Hospital Department of Microbiology, Virology and Infection Prevention & Control we offer the UK’s first UKAS-accredited metagenomics service for CSFs and tissues, most of which are from patients with encephalitis without a diagnosis.4 With 10 years’ experience we have enabled the diagnosis of several cases of encephalitis,5–8 most with rare or unexpected organisms that, if it weren’t for metagenomics, would remain undiagnosed.
References available on our website.


